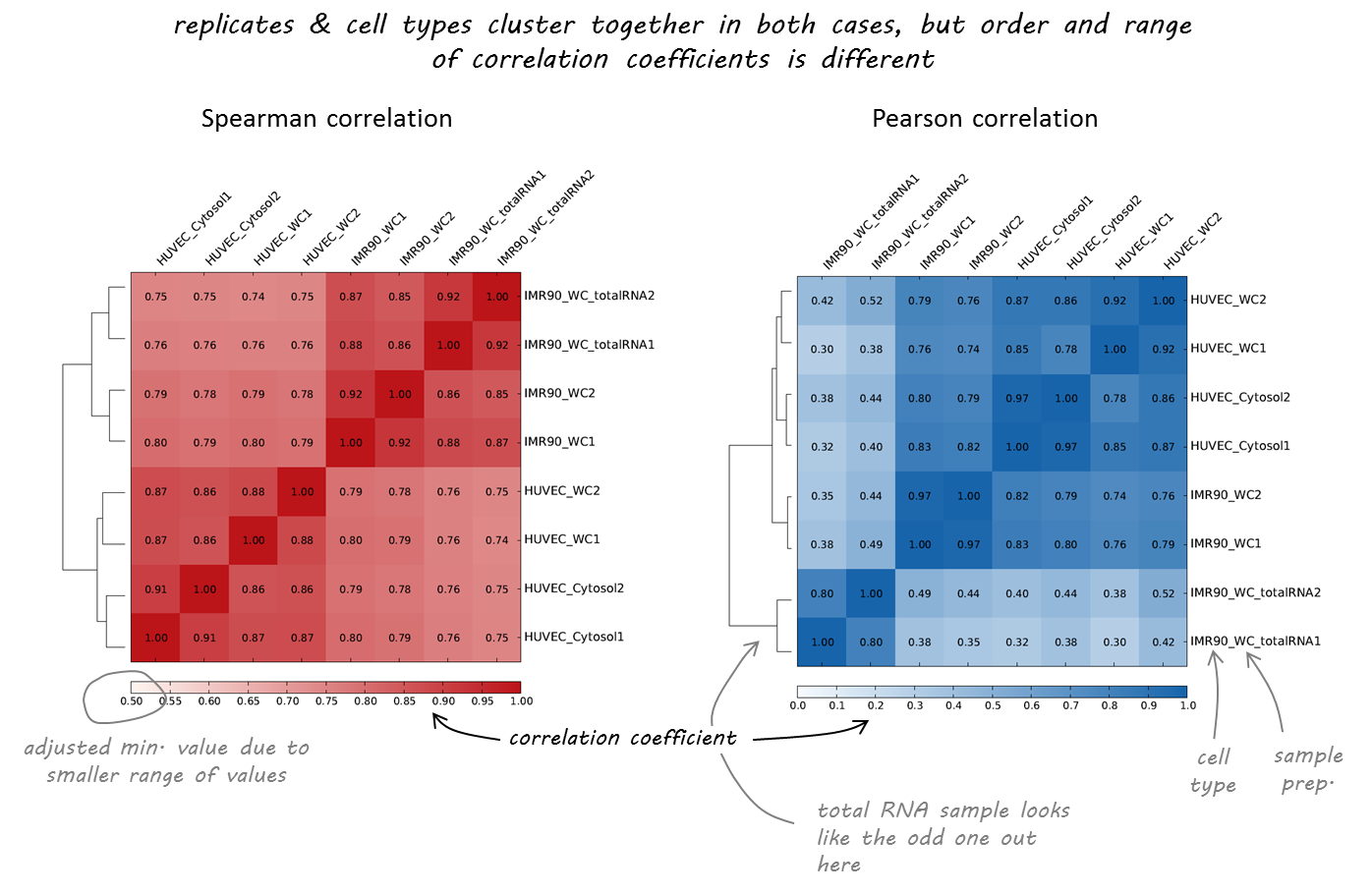
  Type in the gene name "Galnt7" and query its surrounding 400kb. Here we will use the PLAC-Seq (a datatype similar to ChIA-PET) to show chromatin interactions surrounding Galnt7 in mESC as an example.įrom dropdown menu, choose PLAC-Seq, mouse, mm9, mESC, H3K27ac-mm9. 
The main difference for visualizing Hi-C and ChIA-PET/Capture Hi-C is that Hi-C is usually displayed as a heatmap, while the other format are shown as arcs. Visualizing ChIA-PET or Capture Hi-C data By integrating multiple lines of evidence, our browser provides a valuable resource for investigators to create hypotheses connecting distal non-coding variant, distal regulatory elements and their target gene. In this case, virtual 4C, ChIA-PET and DHS-Linkage all support that there is the potential interaction between the enhancer harboring this SNP and the promoter region of PSMA5, which is implicated in heart disease20. This SNP is located within a candidate enhancer region, marked by H3K27Ac as shown in the UCSC genome browser. In the below figure, we show an example by querying rs12740374, a SNPs that has been associated with coronary heart disease19. We incorporated these three methods in our browser. For more details, please check out ENCODE Consortium Nature 2012 and Thurman et al, Nature 2012. Between each pair of distal-proximal DHSs, you can compute a Pearson correlation based their tissue-specificity. Each pairwise ChIA-PET interaction is visualized as elliptical arcs in our browser.ĮNCODE Consortium profiled DHS in more than 100 cell types, and therefore making it possible to compute Pearson correlation coefficients for all distal DHS with gene proximal DHS. In our browser, we use the queried region as bait (either a gene name or a SNP ID), and extract a row of Hi-C data centered on the bait region, hence, virtual 4C.ĬhIA-PET: another implementation of chromatin ligation-based method, which detects long-range interactions between genomic regions that are enriched for a feature (either histone modification or transcription factor binding). 
4C data is plotted as a curve line, where the center is the bait region and a peak signal in distal region indicates there is a chromatin interaction event. Virtual 4C: Circular chromosomal conformation capture(4C) is a chromatin ligation-based method that surveys for one-vs-many interactions in the genome, that is, to measure the interaction frequencies between a bait locus of interest and any other loci. To facilitate users with this goal, we implemented the following three methods in our 3D Genome browser: For example, many users are interested in using Hi-C data to explore enhancer-promoter interactions. Identify Linkage between Genes, Enhancers and SNPs Although displaying Hi-C data as a heatmap is informative to visualize large genome structures such as TADs, it is not intuitive to show interactions between two specific loci. Simply click the "Check gene expression on the top right corner of this page. You can also conveniently check the expression for the queried the genes across over 100 cell types profiled by the ENCODE project. We observe that this known enhancer and the promoter of SHH are located within the same TAD and the long range interaction between them is also evident in the Hi-C map, marked by the black arrow In this region, there is a known enhancer (marked by green bar) that regulate the SHH gene17. In this case, we display the tracks for RNA-Seq and H3K27ac (active enhancer mark) ChIP-Seq data for the same cell type. Under the heatmap, we also imbed the UCSC genome browser for the same region, so that the users can explore both the chromatin interaction and other "omics" data simultaneously. To make the Hi-C interaction more visible, users can conveniently adjust the color bar on the same page. Once the initial HiC heatmap is displayed, there are several other function buttons to further assist users to explore around this region, such as zoom in/out and move to right/left. In Figure below, we display the 25kb resolution Hi-C matrix, published in Rao et al Cell 2014. For Hi-C data with multiple resolutions (bin sizes), there is another dropdown menu for users to choose the appropriate size according to their needs. Our browser auto-fills gene names as users type, based a variety of gene annotations, including refSeq, UCSC and ensemble gene sets.Īfter clicking the submit button, the Hi-C interactions in GM12878 for the queried gene are shown on the right panel. We can visualize this region by selecting human and hg19 from the drop-down for species and genome assembly, and then typing in the gene name SHH in the textbox. #UCSC CPLOT HOW TO#Here we demonstrate how to query the chromatin interactions for regions surrounding SHH (Sonic Hedgehog) gene. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
